2023 Blog, BOTs Blog, RLCatalyst Blog, Blog, Featured
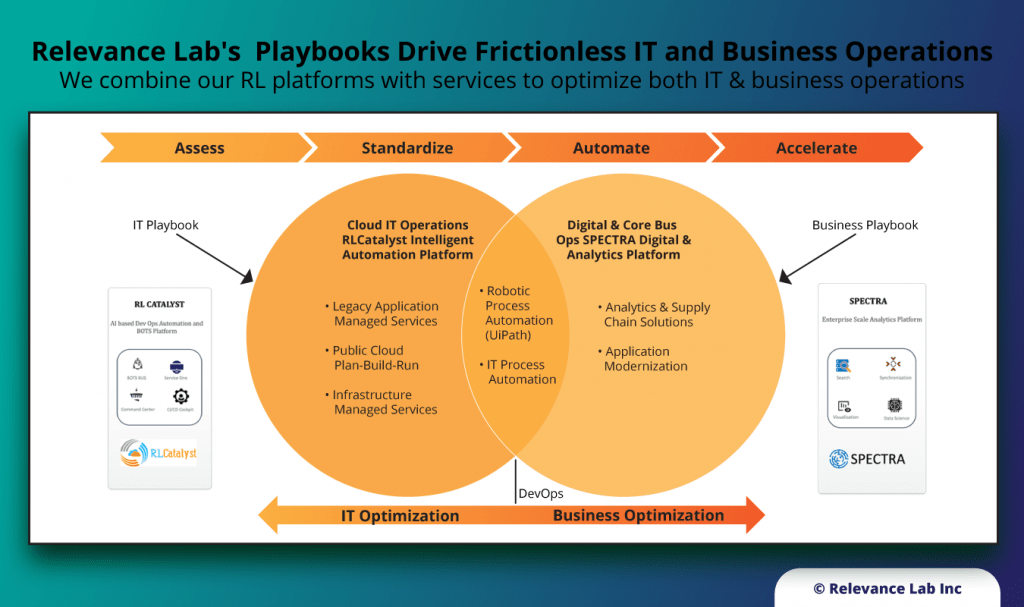
With growing interest & investments in new concepts like Automation and Artificial Intelligence, the common dilemma for enterprises is how to scale these for significant impacts to their relevant context. It is easy to do a small proof of concept but much harder to make broader impacts across the landscape of Hybrid Infrastructure, Applications and Service Delivery models. Even more complex is Organizational Change Management for underlying processes, culture and “Way of Working”. There is no “Silver bullet” or “cookie-cutter” approach that can give radical changes, but it requires an investment in a roadmap of changes across People, Process and Technology. RLCatalyst solution from Relevance Lab provides an Open Architecture approach to interconnect various systems, applications, and processes like the “Enterprise Service Bus” model.
What is Intelligent Automation?
The key building blocks of automation depend on the concept of BOTs. So, what are BOTs?
- BOTs are automation codes managed by ASB orchestration
- Infrastructure creation, updation, deletion
- Application deployment lifecycle
- Operational services, tasks, and workflows – Check, Act, Sensors
- Interacting with Cloud and On-prem systems with integration adapters in a secure and auditable manner
- Targeting any repetitive Operations tasks managed by humans that are frequent, complex (time-consuming), security/compliance related
- What are types of BOTs?
- Templates – CloudFormation, Terraform, Azure Resource Models, Service Catalog
- Lambda functions, Scripts (PowerShell/python/shell scripts)
- Chef/Puppet/Ansible configuration tools – Playbooks, Cookbooks, etc.
- API Functions (local and remote invocation capability)
- Workflows and state management
- UIBOTs (with UiPath, etc.) and un-assisted non-UI BOTs
- Custom orchestration layer with integration to Self-Service Portals and API Invocation
- Governance BOTs with guardrails – preventive and corrective
- What do BOTs have?
- Infra as a code stored in source code configuration (GitHub, etc.)
- Separation of Logic and Data
- Managed Lifecycle (BOTs Manager and BOTs Executors) for lifecycle support and error handling
- Intelligent Orchestration – Task, workflow, decisioning, AI/ML
To deploy BOTs across the enterprise and benefit from more sophisticated automation leveraging AI (Artificial Intelligence), RLCatalyst provides a prescriptive path to maturity as explained in the figure below.
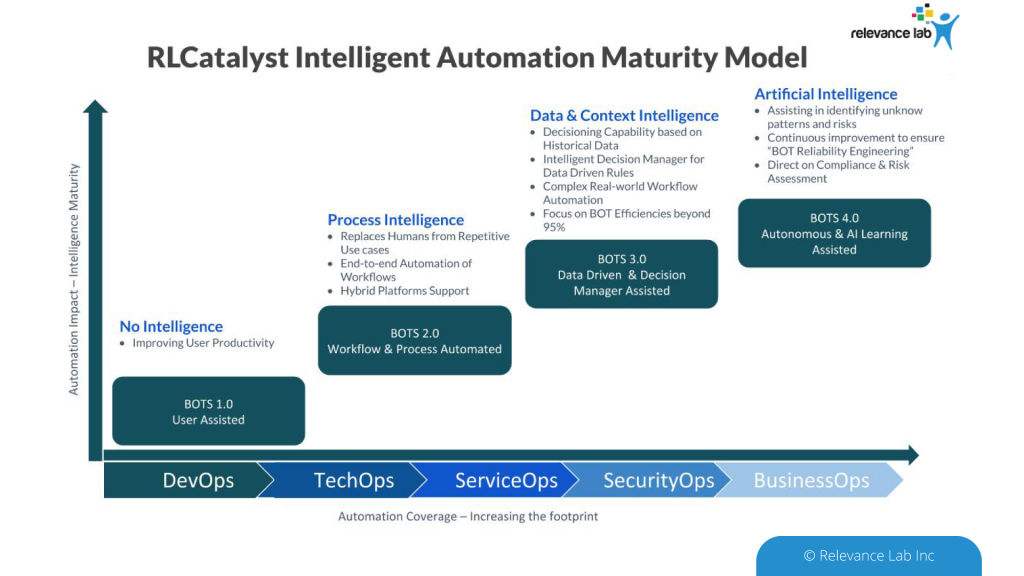
ASB Approach
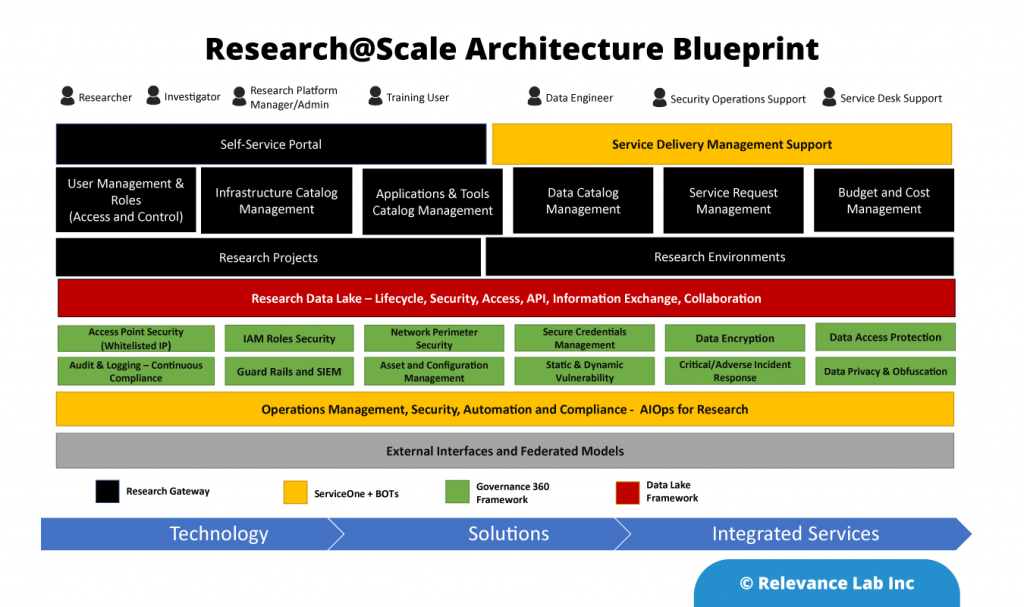
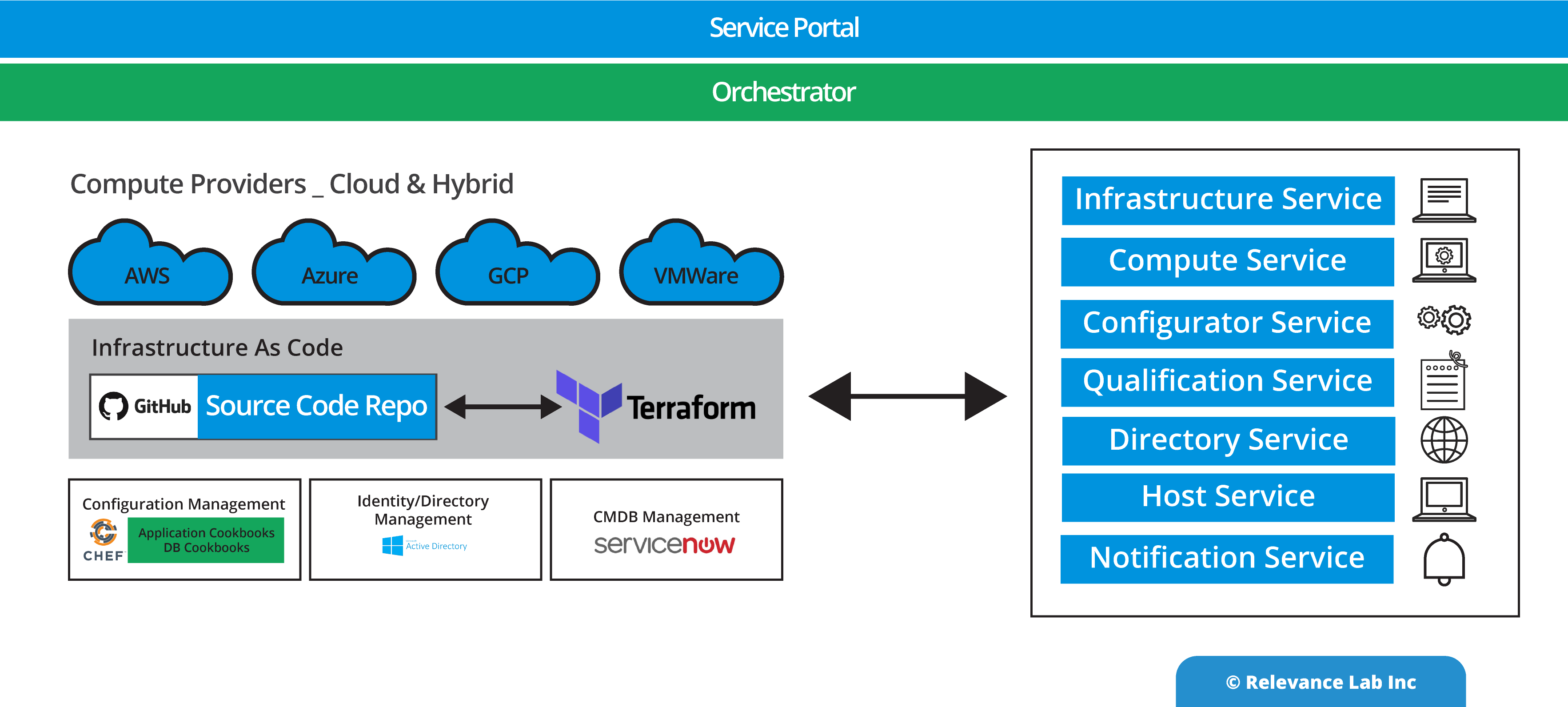
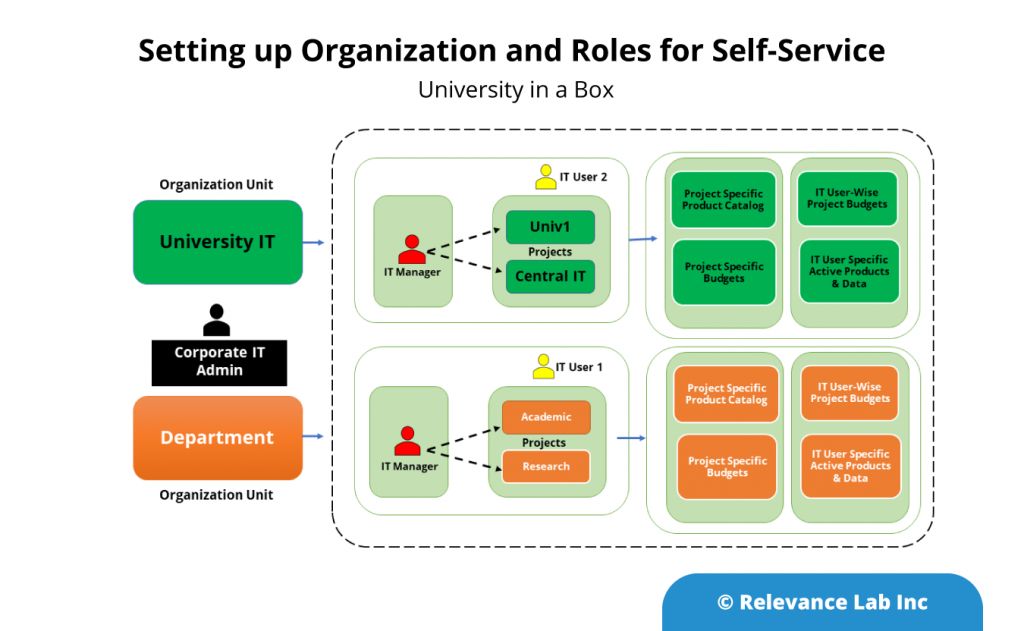
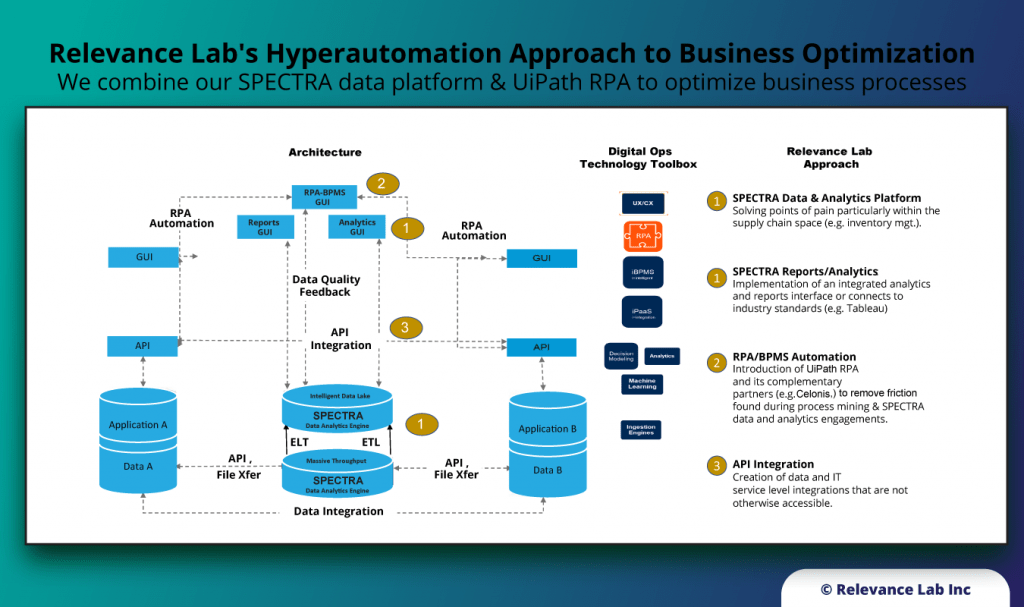
An Open- Architecture approach to interconnect various systems, applications, and processes similar to the “Enterprise Service Bus” model. This innovative approach of “software-defined” models, extendable meta-data for configurations, and a hybrid architecture takes into consideration modern distributed security needs. This ASB model helps to drive “Touchless Automation” with pre-built components and rapid adoption by existing enterprises.
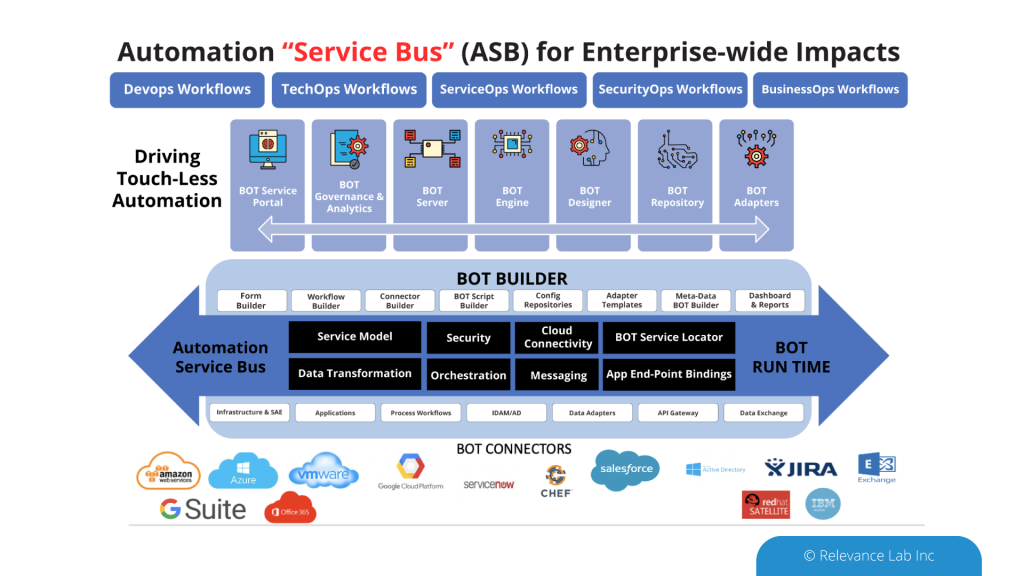
To support a flexible deployment model that integrates with current SAAS (Software as a Service) based ITSM Platforms allows Automation to be managed securely inside Cloud or On-Premise data centers. The architecture supports a hybrid approach with multi-tenant components along with secure per instance-based BOT servers managing local security credentials. This comprehensive approach helps to scale Automation from silos to enterprise-wide benefits of human effort savings, faster velocity, better compliance and learning models for BOT efficiency improvements.
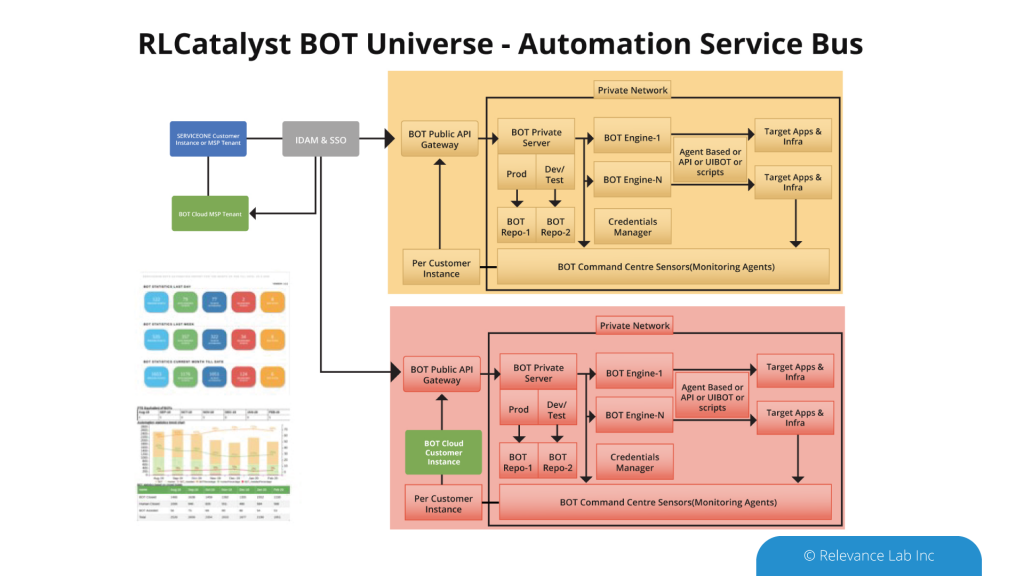
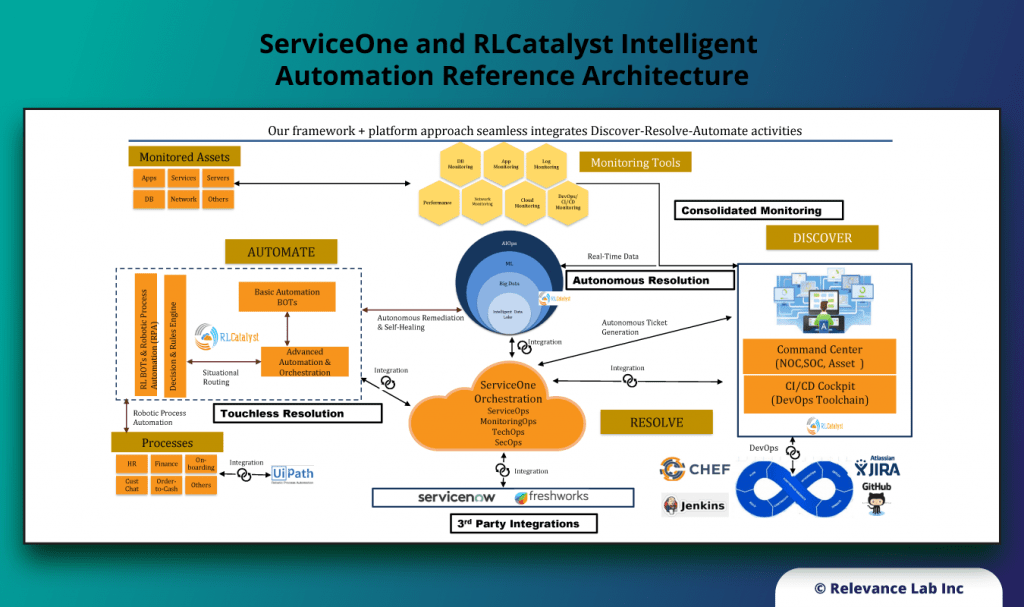
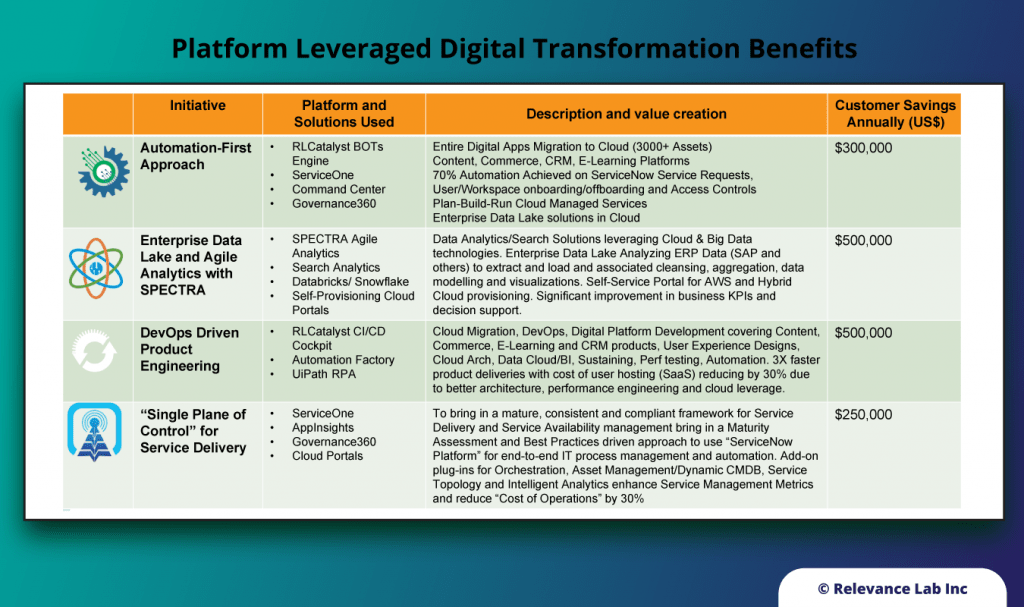
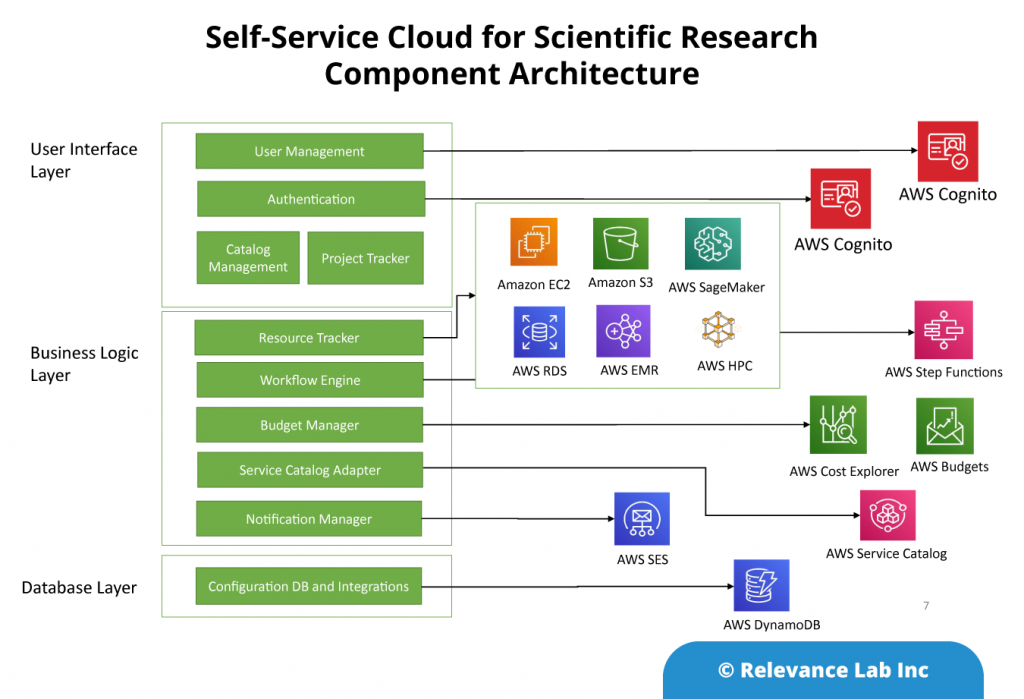
RLCatalyst provides solutions for enterprises to create their version of an Open Architecture based AIOps Platform that can integrate with their existing landscape and provide a roadmap for maturity.
- RLCatalyst Command Centre “Integrates” with different monitoring solutions to create an Observe capability
- RLCatalyst ServiceOne “Integrates” with ITSM solutions (ServiceNow and Freshdesk) for the Engage functionality
- RLCatalyst BOTs Engine “Provides” a mature solution to “Design, Run, Orchestrate & Insights” for Act functionality
Relevance Lab is working closely with leading enterprises from different verticals of Digital Learning, Health Sciences & Financial Asset Management in creating a common “Open Platform” that helps bring Automation-First approach and a maturity model to incrementally make Automation more “Intelligent”.
For more information feel free to contact marketing@relevancelab.com
References
Get Started with Building Your Automation Factory for Cloud
Intelligent Automation For User And Workspace Onboarding
Intelligent Automation with AS/400 based Legacy Systems support using UiPath
RLCatalyst BOTs Service Management connector for ServiceNow